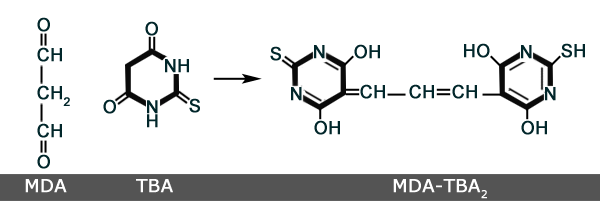
The Trouble with TBARS
(and how to avoid it)
360-449-3091
Mentioned in connection with oxidized lipids as early as 1948(1) Thiobarbituric Acid Reactive Substances (TBARS) are often mentioned in discussions about malondialdehyde (MDA). However, it is important to understand that MDA is one of many substances that can and do react with TBA. Therefore, as the name implies, the TBARS assay is not totally specific for MDA(2). However, since MDA is one of the less reactive(3) aldehydes, it is generally considered to account for almost all of the color development in the TBARS reaction. Despite calls for caution(2), many researchers have historically used the assay as a simple, cost effective surrogate for more specific methods of MDA testing which are generally more labor intensive.
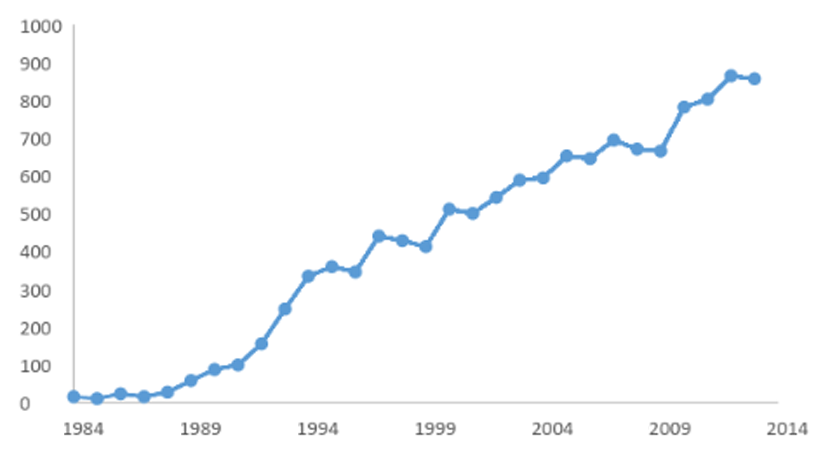
Acceptance and Persistence of TBARs in Scientific Publications
(4)
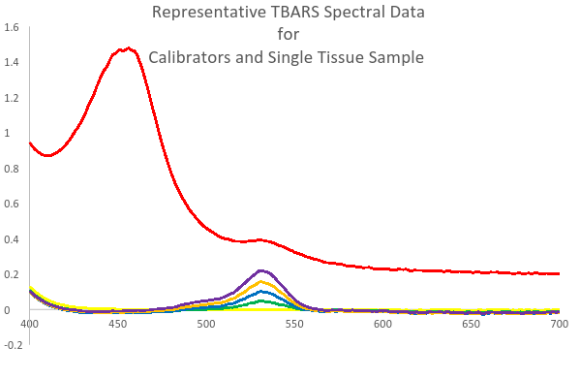
Users of the TBARS assay should be aware that all samples with complex protein content will produce a spectral pattern and baseline shift very different than the spectral pattern observed for calibrators in a pure system.
As can be seen, calibrators run in a pure system produce a normal zero baseline whereas the complexities in the tissue sample matrix yield an elevated and nonlinear baseline. Because of this shifted and skewed baseline, a straight up comparison of uncorrected, 532 nm sample TBARS data with calibrator peak data is impossible. Doing so, produces inherently erroneous results and sample to sample comparisons can even be problematic without some sort of blanking or analysis based correction being applied.
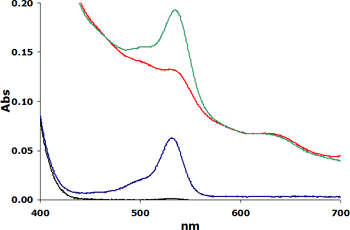
Will extraction alone deal with issue?
Does this mean that all TBARS data is incorrect?
No. Fortunately, there are ways to correct for the inherent “trouble with TBARs”. We recommend that investigators pay close attention to how TBARS data was generated and whether any corrective measures were employed in the analysis of sample data.
Two such corrective measures are as follows:
- Subtract the value at 572 nm where the curve is presumed to begin (532-572)
- Collect full “scan” data from 400-700 nm and perform derivative analysis to quantify only the true area under the curve of interest
An in depth discussion of various analysis strategies for the MDA-TBA2 chromophore can be found under resources for our NWK-MDA01 assay:
Additionally, we provide an online derivative analysis program that allows researchers to upload 400-700 nm scan data for calibrators and samples in order to generate the best possible TBARS generated data.
Assuming researchers take the time to follow one of the above methods, TBARS based assays can and do produce excellent data regarding levels of lipid peroxidation in biological samples.
References:
- Bernheim F, Bernmeim ML, Wilbur KM. The reaction between thiobarbituric acid and the oxidation products of certain lipides. J Biol Chem. 1948 May;174(1):257-64.
- Janero, D.R. Malondialdehyde and thiobarbituric acid-reactivity as diagnostic indices of lipid peroxidation and peroxidative tissue injury, Free Rad. Biol. Med. 9:515-540; 1990.
- Esterbauer, H., et.al., Chemistry and Biochemistry of 4-Hydroxynonenal, malondialdehyde and Related Aldehydes, Free Rad. Biol. Med. 11:81-128; 1991.
- PubMed: Results by Year: (https://www.ncbi.nlm.nih.gov/pubmed/?term=TBARS)
- Jentzsch, A. M., et.al., Improved Analysis of Malondialdehyde in Human Body Fluids, Free Rad. Biol. Med. 20:251-256; 1996.
- Botsoglou, N.A., Rapid, Sensitive, and Specific Thiobarbituric Acid Method for Measuring Lipid Peroxidation in animal Tissue, Food and Feedstuff Samples, J. Agric. Food Chem. 42:1931-1937; 1994.
Malondialdehyde (MDA) Assay Kit: 200 Tests
A colormetric assay kit for detection of Malondialdehyde (MDA) or Thiobarbituric Acid Reactive Substances (TBARS) in multiple species and sample types. Designed for simplicity and affordability, this assay utilizes various improvements to provide you with more dependable data.
Advantages of the NWLSS™ Malondialdehyde Assay:- BHT and EDTA are added to the sample and reaction mixture to minimize artifact oxidation.
- Reduced reaction temperature minimizes the decomposition of lipid hydroperoxides.
- Optimized reaction pH facilitates hydrolysis of MDA-protein adducts for better recovery of malondialdehyde (MDA).
- Cleaner output through optimized data reduction using dual wavelength or 3rd derivative analysis of spectroscopic scan data.